|
Background for
Bioinformatics Stats Investigation |
|
|
How genetic information is stored: The
information stored in DNA is remarkably similar to a low level computer language
written with 4 letters:
A,
C,
G,
T. Each of
these represents a different base arranged in a strand of DNA. A pair of
complementary strands are joined with hydrogen bonds and form the famous double
helix.
DNA Sequences: To sequence DNA, the bases are
read from a single strand a few pieces at a time and then assembled into a
lengthy string of As, T, G, and Cs. An entire set of these strings for a single
organism is called a Genome.
|
|
| The human
genome contains 3,164.7 million of these letters as compare to 3.6
million letters in the King James Version of the Bible. The letters in
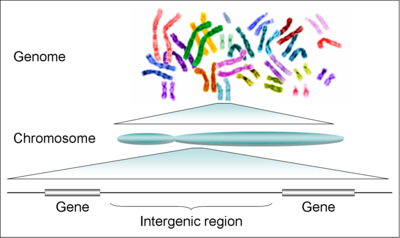
the genome are arranged like the chapters in a book on 24 separate DNA
molecules each contained in a different chromosome. Each chromosome
contains numerous genes which are considered the basic functional
units of heredity.
Genes vs. intergenetic regions:
Although the human genome is thought to contain about 20,000 to 25,000
genes, the genes make up only about 2% of the human genome. The
remaining 98% of DNA is called the intergenetic regions. It purpose is
unknown, although, even a relatively simple analysis indicates that it
contains some form of information--maybe useless, scrambled or
abandoned information.
|
-
genomepunctuationquitesimplythereisntanytherearenosp
-
acesnopunctuationmarksorcapitalletterstodenotethebegi
-
nningsandendingsofwordsinasequencetryreadingtheparag
-
raphatrightforacomparisonofhowthiswouldlookinenglish
|
|
Genome punctuation: Quite simply there isn't
any. There are no spaces, no punctuation marks, or capital letters to denote the
beginnings and endings of words in a sequence. Try reading the paragraph at
right for a comparison of how this would look in English.
|
|
Statistical analysis of genomes: Statistics is probably the single
most important mathematical tool for understanding genetics. The
investigation below will help show some of the many ways it can be
used. |
| |
|
Stats
Investigation: Statistical Analysis of Genome Data - time approx 2 class periods |
|
Purpose:
Determine if a statistical method can determine
whether a sequence is randomly generated or contains actual information. Instructions
- Begin by comparing the two
sequences shown in the link.
One is a real genetic sequence, the other a randomly
generated sequence. Can you tell the difference just by
looking?
- Go the article about
Genome mining, "Meaningful
sequences" and read through it to the end. (You will
have to click on the small arrow icon on the right side
below the text.)
- Now go to "Analyzing
DNA" and follow the exercises using Geneboy all the
way to the end of the article.
- Using Analyze composition,
Singles feature of Geneboy obtain the distribution of
frequencies for A, C, G, T bases for the following 3
cases: 1) Genetic 1, 2) Random 1, and 3) Intergenetic 1. What type of distribution would
be predicted using the law of large numbers for a random
distribution of As, Cs, Gs, and Ts? What statistical tool
could you use to confirm whether the distributions are or
are not random? Perform this analysis on for all 3 cases. Note, the question
is not asking if the two distributions are different from
each other.
- Plot 2 bar graphs using
the distribution of frequencies for A, C, G, and T of
Genetic 1. In the first scale the y-axis from 15 to 35%,
the second from 0 to 300%. Repeat the process for Random
1.
Questions /Conclusions:
- Did the changing the scale
of the above graphs alter the perception of their meaning
and could it alter the meaning of a different set of
graphs?
- Which is more likely to be
universally accepted, conclusions based on graphs or
conclusions based on statistical analysis? Why?
- If genes are only 2% of
the sequence how would you find one and establish that it
was one using statistics?
|
|
|